Detect Neutralizing Antibodies Against Omicron and Other SARS-CoV-2 Variants
Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) is the infectious agent responsible for the worldwide Coronavirus Disease 2019 (COVID-19) pandemic1. As the virus replicated and spread globally, its genome sequences underwent alterations, referred to as mutations, and the virus continued to evolve. It became clear in the early phase of the pandemic that SARS-CoV-2 evolution would be a major issue2. The speed by which the SARS-CoV-2 virus changed, in turn, drove the need for advanced technologies in diagnostic tests and mutation surveillance to keep pace. Our ability to detect antibodies generated against this virus is also of paramount importance. In this blog, we highlight our advanced solution to measure anti-SARS-CoV-2 antibodies, critical to help evaluate the immune status of the population as we face this rapidly evolving virus together.
A variant is defined as a population of coronaviruses that inherit the same distinctive mutations. While scientific nomenclature for naming SARS-CoV-2 lineages is advantageous for researchers, broader discussions of lineages have been facilitated by nomenclature put forth by the WHO2 and will be utilized here. Classification of variants of SARS-CoV-2 includes variant of interest (VOI), variant of concern (VOC), and variant under monitoring (VUM) (see side bar for definitions).
By August 30, 2021, four VOCs (Alpha, Beta, Gamma, and Delta) were designated by the WHO3. More recently, two VOCs (Delta and Omicron) are recognized (updated April 25, 2022)2. Emergence of these variants is primarily driven by mutations arising in the spike protein, impacting protein structure. This leads to modified interaction with the human ACE2 receptor and thereby infection rates4. To date, the Omicron variant is the most divergent strain we have experienced during the pandemic (Figure 1)4, with changes being linked to greater transmissibility (Figure 2).
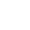
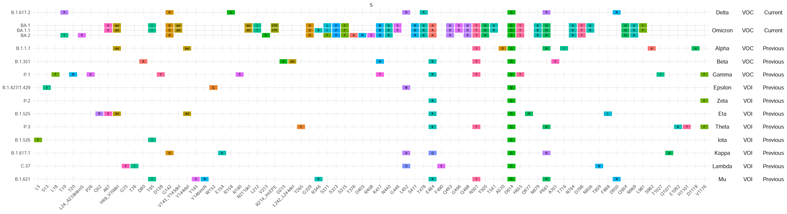
Figure 1. VOI/VOC profiles of Spike amino acid changes. For each variant, the profile of amino acid changes compared to the wild-type virus (GISAID Accession ID: EPI_ISL_402124) was created based on the first 1,000 genomes (for Theta, it is based on 544 genomes) available in GISAID (genomes with less than 29,000 nucleotides and >5% Ns were excluded). Amino acid changes that are present in ≥ 85% of the analyzed sequences are shown. Relevant amino acid changes may be present in other regions of the SARS-CoV-2 genome. Not all amino acid changes in the spike protein are associated with potential changes in the characteristics of the virus variant2.
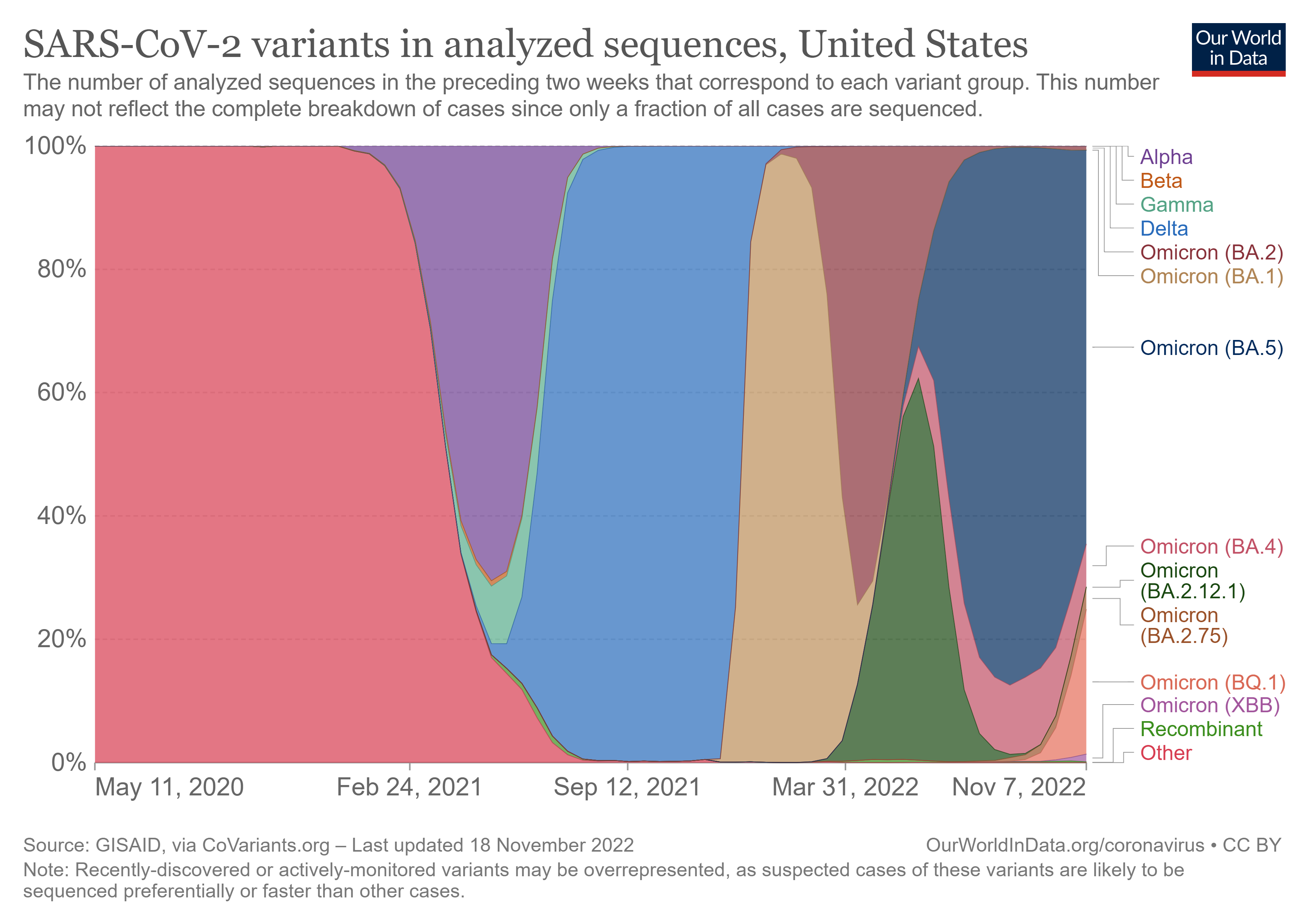
Figure 2. SARS-CoV-2 variants in analyzed sequences, United States. The number of analyzed sequences in the preceding two weeks that correspond to each variant group. This number may not reflect the complete breakdown of cases since only a fraction of all cases are sequenced5.
Now that we know the complexity and speed by which the SARS-CoV-2 virus changes, technologies to detect antibodies generated against this virus must keep pace. Assessment of neutralizing antibodies (NAbs) is of particular importance because these are the antibodies that inhibit the binding of the spike RBD to the human ACE2 receptor, thereby blocking virus entry into host cells and possibly reducing infection and disease severity. Assays that reliably measure NAbs can assess true protective immunity of antibodies and, subsequently, the immune status of our population.
Our LEGENDplex™ SARS-CoV-2 Variants Neut. Ab Panel (6-plex) accurately measures neutralizing antibodies against six SARS-CoV-2 spike protein variants: Alpha, Beta, Gamma, Wild type S1, Delta, and Omicron (Figures 3 and 4). We also offer options for researchers who may need to assess NAbs to a specific variant or a subset of variants. Our Mix & Match panel builder allows you to select variant-specific capture beads and plate format desired. Choose SARS-CoV-2 Variants Neutralization Antibody Panel (6-plex) to view full selection. Trust our reagents to advance your research.
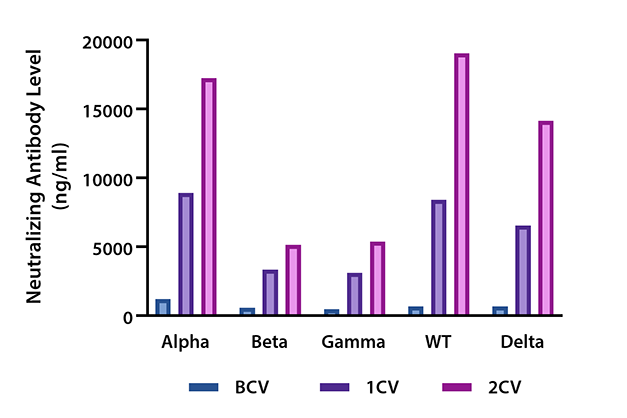
Figure 3. Serum samples from 32 donors were collected before vaccination (BCV), 2 weeks after the 1st dose of vaccination (1CV), and 2 weeks after the 2nd dose of vaccination (2CV) (Moderna or Pfizer SARS-CoV-2 vaccines) then tested for variants neutralization antibody levels against SARS-CoV-2 Spike protein Variants Alpha (B.1.1.7), Beta (B.1.351), Gamma (P.1), Wild type S1, and Delta (B.1.617.2) using the LEGENDplex SARS-CoV-2 Variants Neutralization Antibody Panel (5-plex).
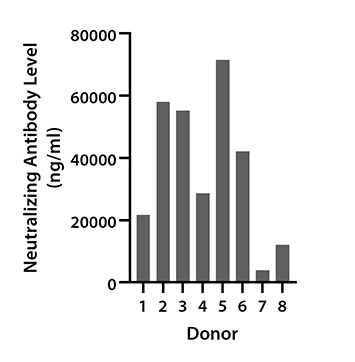
Figure 4. Serum samples from 8 donors were collected between June and September 2022 and tested for neutralization antibody concentrations against SARS-CoV-2 Spike protein Variant Omicron using LEGENDplex SARS-CoV-2 Variants Neut. Ab Panel (6-plex). Vaccination status of donors was not disclosed.
Streamlining the workflow to measure antibodies raised against SARS-CoV-2 infection or in response to vaccination helps us track the status of protective immunity in our population and is critical to our COVID-19 response. At BioLegend, our scientists have designed immunoassays for the specific and sensitive quantitation of anti-SARS-CoV-2 antibodies. Armed with such top-quality research, we all move toward a global vaccination strategy and life after COVID-19.
Resources:
- Wu, Fan et al. “A new coronavirus associated with human respiratory disease in China.” Nature vol. 579,7798 (2020): 265-269. doi:10.1038/s41586-020-2008-3. PubMed
- “Tracking Sars-COV-2 Variants.” World Health Organization, 2022, https://www.who.int/en/activities/tracking-SARS-CoV-2-variants
- Choi, Jun Yong, and Davey M Smith. “SARS-CoV-2 Variants of Concern.” Yonsei medical journal vol. 62,11 (2021): 961-968. doi:10.3349/ymj.2021.62.11.961. PubMed
- Kumar, Suresh et al. “Omicron and Delta variant of SARS-CoV-2: A comparative computational study of spike protein.” Journal of medical virology vol. 94,4 (2022): 1641-1649. doi:10.1002/jmv.27526. PubMed
- “SARS-COV-2 Variants in Analyzed Sequences.” Our World in Data, 2022, https://ourworldindata.org/grapher/covid-variants-area?country=~USA
 Login/Register
Login/Register 






Follow Us